Connecting Databricks to CluedIn
January 14, 2024
In the previous article, we loaded data from CluedIn to Databricks with the help of the cluedin library and a Jupyter notebook. In this article, we will do the opposite: load data from Databricks to CluedIn.
Let's start with a simple example. We have a table named imdb_titles in Databricks with 601_222 rows:

CluedIn
We want to load this data to CluedIn. To do that, we need to create an API token in CluedIn. Go to Administration > API Tokens and create a new token:
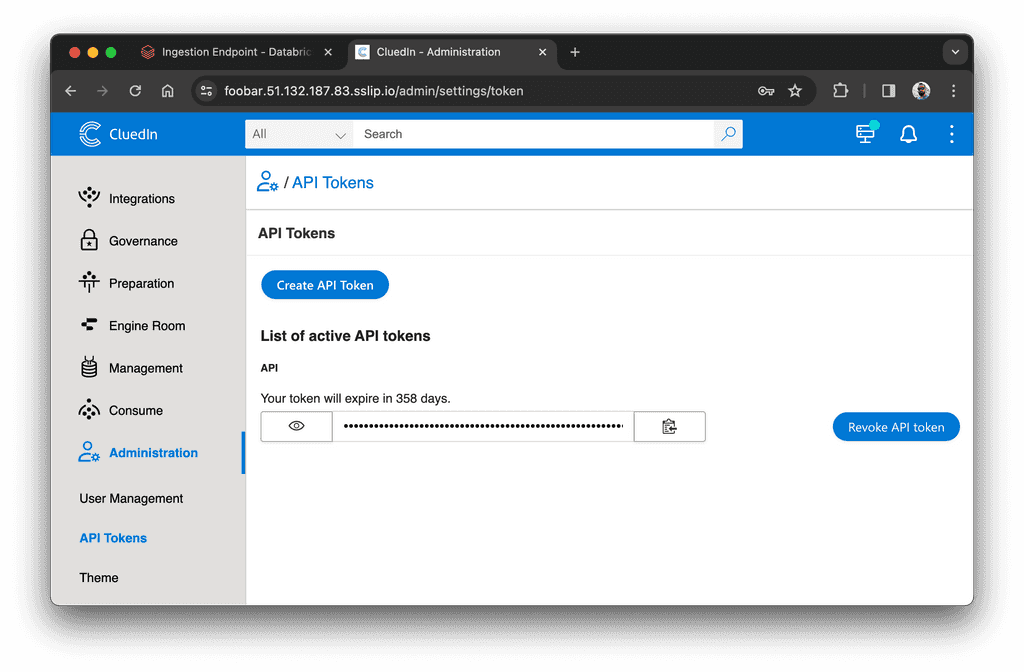
Next, create an endpoint in CluedIn. From CluedIn's main page, click "Import From Ingestion Endpoint" and create a new endpoint. You will need to enter the endpoint's name, group name, and select entity type:
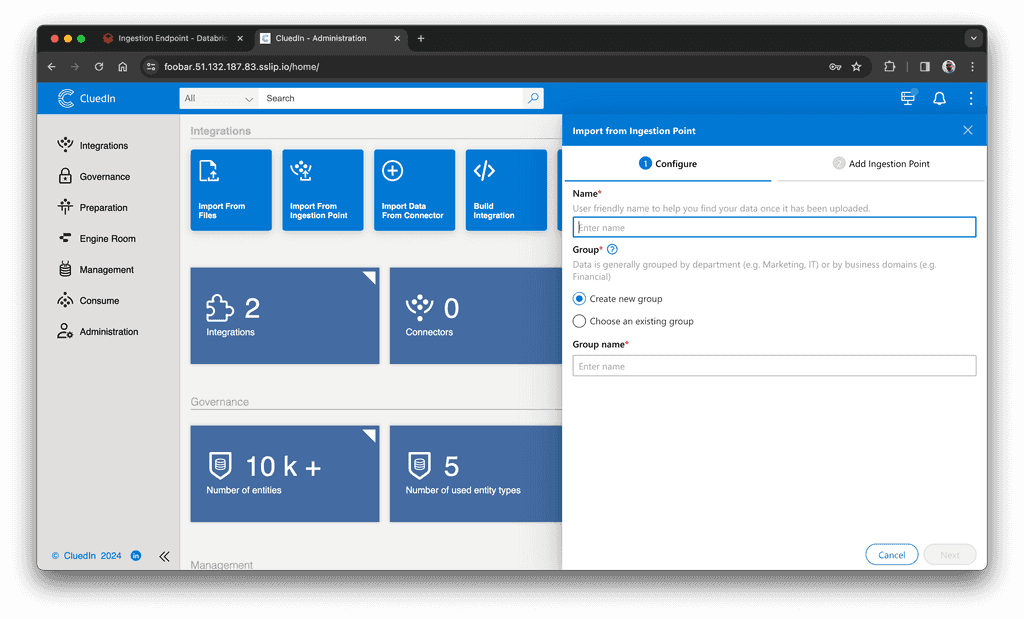
After you create the endpoint, you will find its URL in the "View Instructions" section:
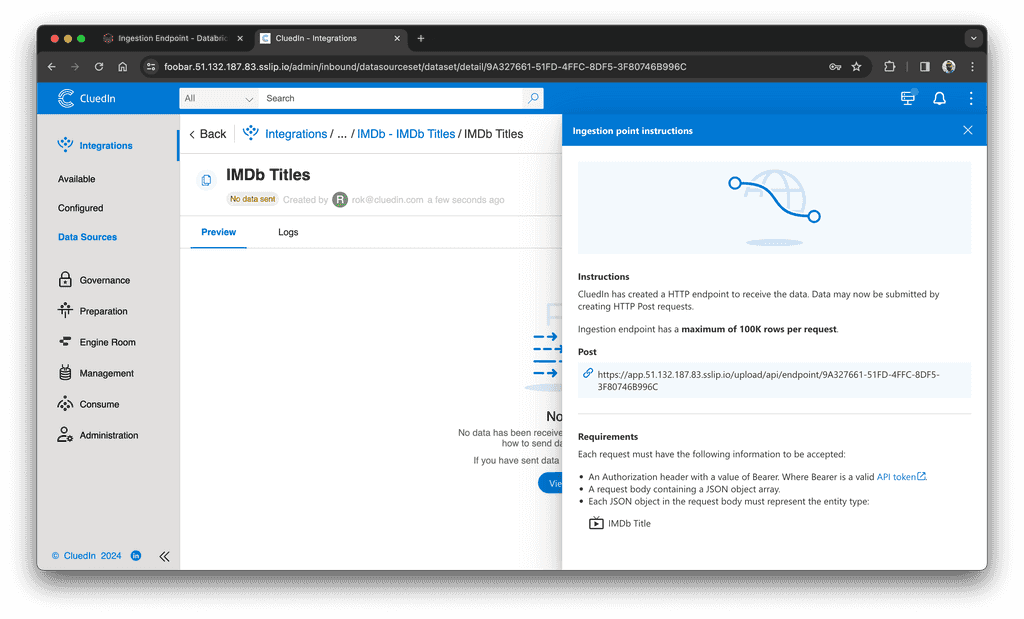
We are ready to load data to CluedIn. Now it's time to set up Databricks.
Databricks
In Databricks, we create a Jupyter notebook and install the cluedin library:
%pip install cluedin
Next, we import the cluedin library and create a CluedIn context object:
import requests
import cluedin
ctx = cluedin.Context.from_dict(
{
"domain": "51.132.187.83.sslip.io",
"org_name": "foobar",
"access_token": "(your token)",
}
)
ENDPOINT_URL = "https://app.51.132.187.83.sslip.io/upload/api/endpoint/9A327661-51FD-4FFC-8DF5-3F80746B996C"
DELAY_SECONDS = 5
BATCH_SIZE = 100_000
In our example, the URL of our CluedIn instance is https://foobar.51.132.187.83.sslip.io/, so the domain is 51.132.187.83.sslip.io and the organization name is foobar. The access token is the one we created earlier.
Next, we will pull data from Databricks and post it to CluedIn.
Here is a simple method to select all rows from a table and yield them one by one:
from pyspark.sql import SparkSession
def get_rows():
spark = SparkSession.builder.getOrCreate()
imdb_names_df = spark.sql("SELECT * FROM hive_metastore.cluedin.imdb_titles")
for row in imdb_names_df.collect():
yield row.asDict()
Next, we create a method that posts a batch of rows to CluedIn:
import time
from datetime import timedelta
def post_batch(ctx, batch):
response = requests.post(
url=ENDPOINT_URL,
json=batch,
headers={"Authorization": f"Bearer {ctx.access_token}"},
timeout=60,
)
time.sleep(DELAY_SECONDS)
return response
def print_response(start, iteration_start, response) -> None:
time_from_start = timedelta(seconds=time.time() - start)
time_from_iteration_start = timedelta(
seconds=time.time() - iteration_start)
time_stamp = f'{time_from_start} {time_from_iteration_start}'
print(f'{time_stamp}: {response.status_code} {response.json()}\n')
print_response is a helper method that prints the response status code and the response body.
Finally, we iterate over the rows and post them to CluedIn. Note that we are posting the rows in batches of BATCH_SIZE rows. DELAY_SECONDS is the number of seconds to wait between batches.
However, we will need to post a small batch of rows first to set up mapping on the CluedIn side. We will post ten rows. To do that, we will add the following lines in the code below:
if i >= 10:
break
Here is the code that posts the rows to CluedIn:
batch = []
batch_count = 0
start = time.time()
iteration_start = start
for i, row in enumerate(get_rows()):
if i >= 10:
break
batch.append(row)
if len(batch) >= BATCH_SIZE:
batch_count += 1
print(f'posting batch #{batch_count:_} ({len(batch):_} rows)')
response = post_batch(ctx, batch)
print_response(start, iteration_start, response)
iteration_start = time.time()
batch = []
if len(batch) > 0:
batch_count += 1
print(f'posting batch #{batch_count:_} ({len(batch):_} rows)')
response = post_batch(ctx, batch)
print_response(start, iteration_start, response)
iteration_start = time.time()
print(f'posted {(i + 1):_} rows')
After we run the code, we shall see ten rows in CluedIn:
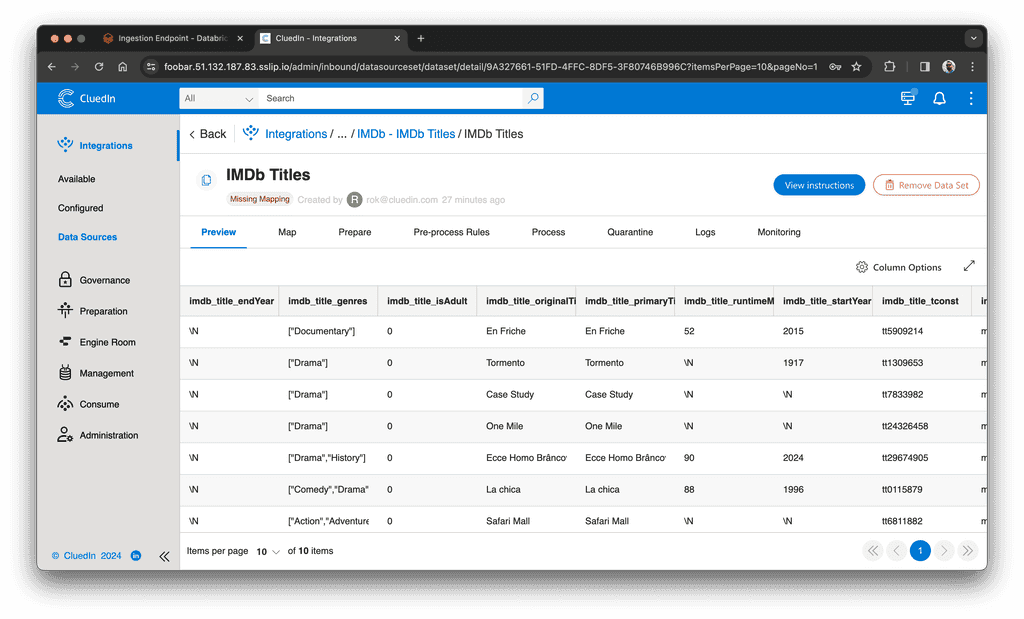
Then, in the Map tab, we create an automatic mapping:
- Click "Add Mapping".
- Select "Auto Mapping" and "Next".
- Ensure the entity type is selected or type a new entity type name and click "Create".
- Type the new vocabulary name, like
imdb.titleand click "Create". - Click "Create Mapping".
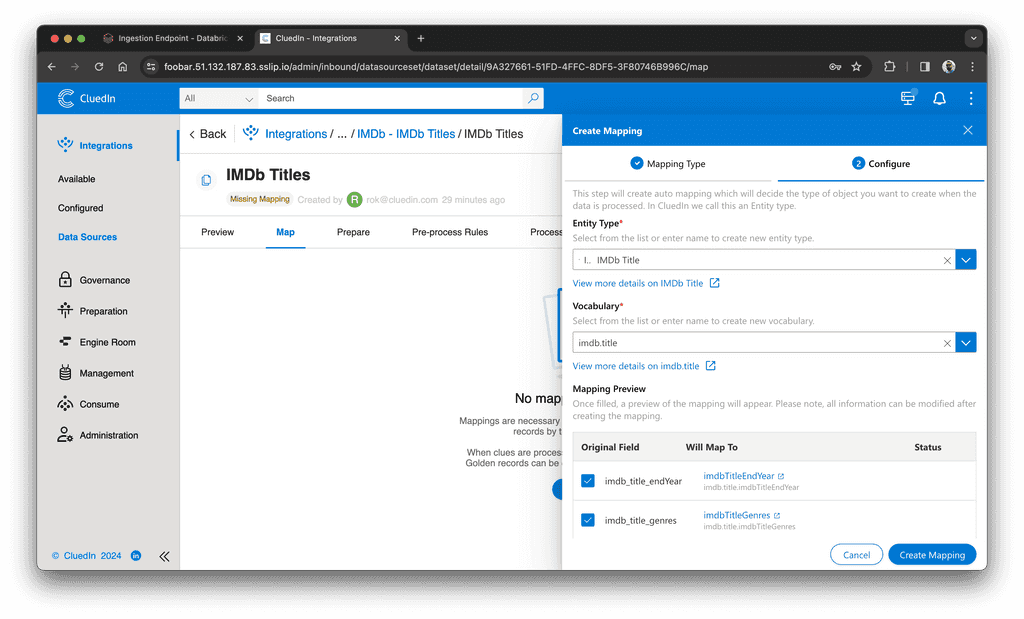
- Click "Edit Mapping".
- On the "Map Entity" tab, select the property used as the entity name and the property used for the origin entity code:
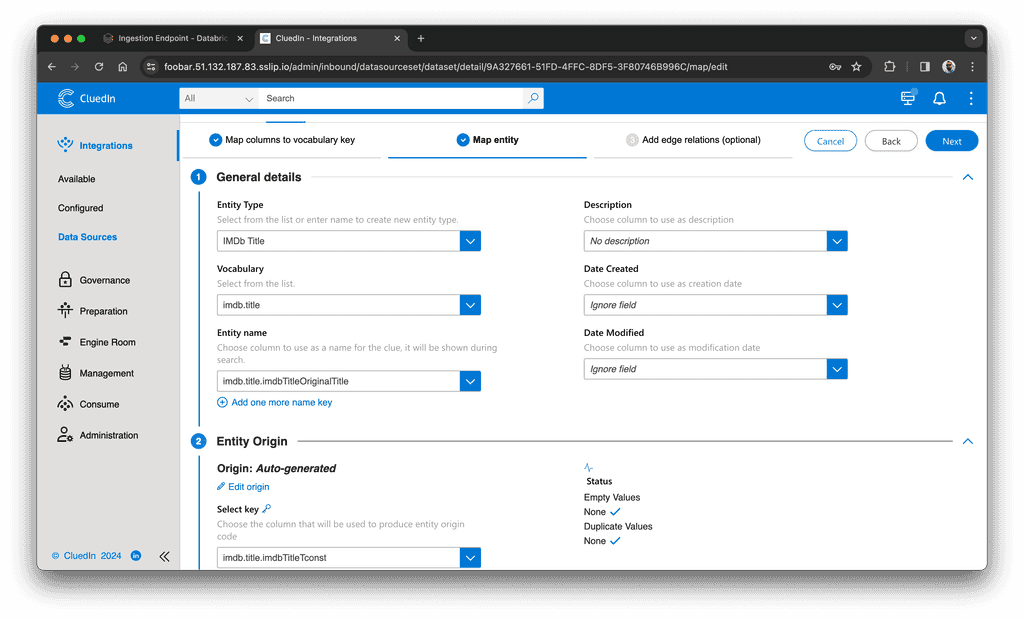
- Click "Next" and click "Finish".
- On the "Process" tab, enable "Auto submission" and then click "Switch to Bridge Mode":
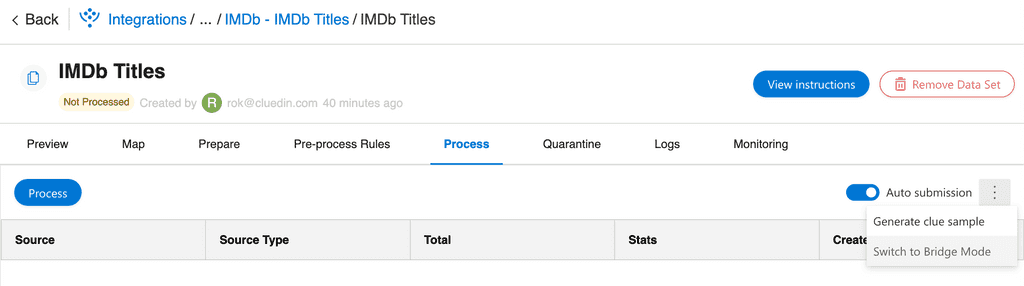
- After you followed the instructions, you can remove or comment on these lines in the notebook and rerun it:
if i >= 10:
break
All table rows will be posted to CluedIn. The output in the notebook will look like this:
0:00:13.398879 0:00:13.398889: 200 {'success': True, 'warning': False, 'error': False}
posting batch #2 (100_000 rows)
0:00:22.021498 0:00:08.622231: 200 {'success': True, 'warning': False, 'error': False}
posting batch #3 (100_000 rows)
0:00:30.709844 0:00:08.687518: 200 {'success': True, 'warning': False, 'error': False}
posting batch #4 (100_000 rows)
0:00:40.026708 0:00:09.316675: 200 {'success': True, 'warning': False, 'error': False}
posting batch #5 (100_000 rows)
0:00:48.530380 0:00:08.503460: 200 {'success': True, 'warning': False, 'error': False}
posting batch #6 (100_000 rows)
0:00:57.116517 0:00:08.585930: 200 {'success': True, 'warning': False, 'error': False}
posting batch #7 (1_222 rows)
0:01:02.769714 0:00:05.652984: 200 {'success': True, 'warning': False, 'error': False}
posted 601_222 rows
Give CluedIn some time to process the data, and you should see 601_222 rows in CluedIn.
The same approach works not only with Databricks but also with any other data source. You must change the get_rows method to pull data from your source.